EDT-Former: Adaptive Connectors Let Frozen LLMs Handle Entities of Any Complexity
ICLR 2026
Situation: Connectors that link structure encoders to LLMs use a fixed number of tokens regardless of entity complexity — losing critical structural detail for large entities while wasting capacity for simple ones, and requiring expensive LLM fine-tuning to compensate. Task: Design a connector that automatically scales its token budget to entity complexity without touching the LLM backbone. Action: Built an entropy-guided dynamic connector that identifies information-dense boundaries in each entity's structure and allocates more tokens where complexity is high, keeping both encoder and LLM fully frozen during training. Result: Achieves top-1 across property prediction, reasoning, and generation benchmarks, running 3.5× faster with half the GPU memory of standard fine-tuning — making scalable structure-to-language alignment practical.
Background & Challenge
Two failure modes in connecting structure encoders to language models:
1. Fixed-budget connectors lose structural detail. Current connectors fix the number of query tokens regardless of entity size. For small entities, a tight budget still captures key features — but for larger ones, the same budget misses critical substructural context that cannot be recovered by prompting alone.
2. LLM fine-tuning is prohibitively expensive. Competitive systems jointly fine-tune the LLM backbone: 8B+ trainable parameters and nearly 3× more compute per token than connector-only training. No prior connector-only method had matched the performance of full fine-tuning under a fully frozen LLM regime.
Methodology
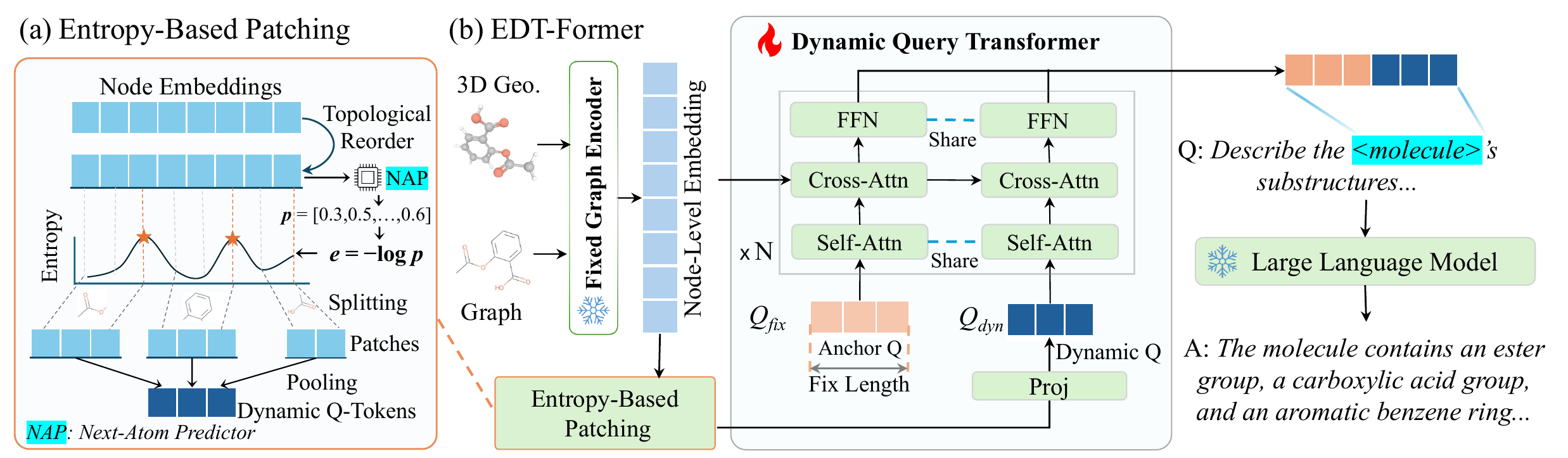
Entropy-Based Boundary Detection
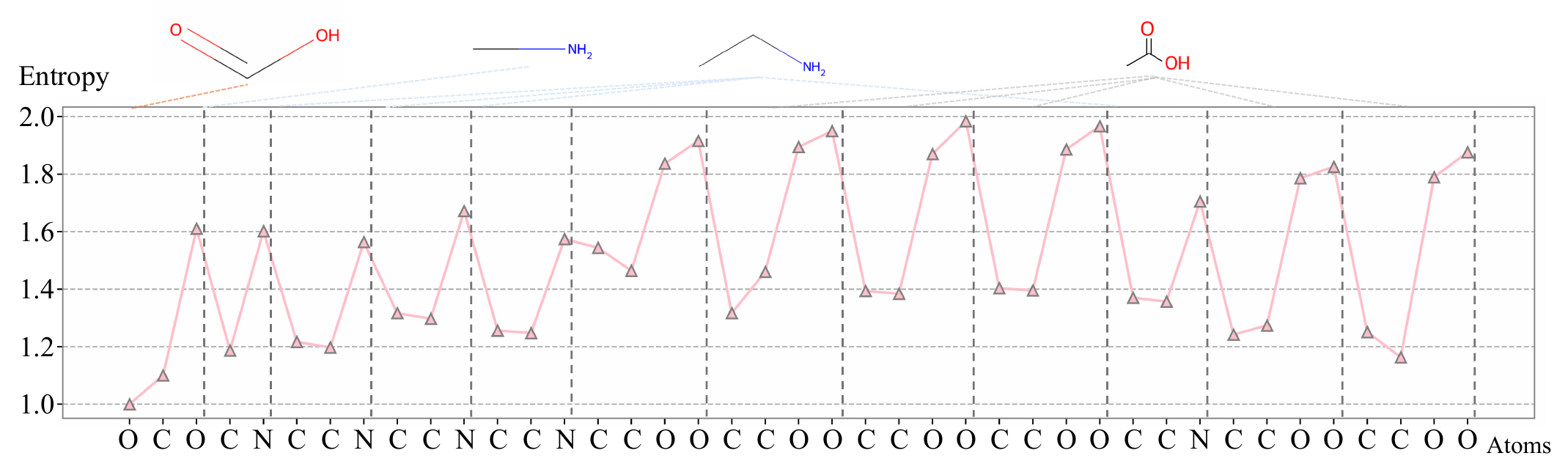
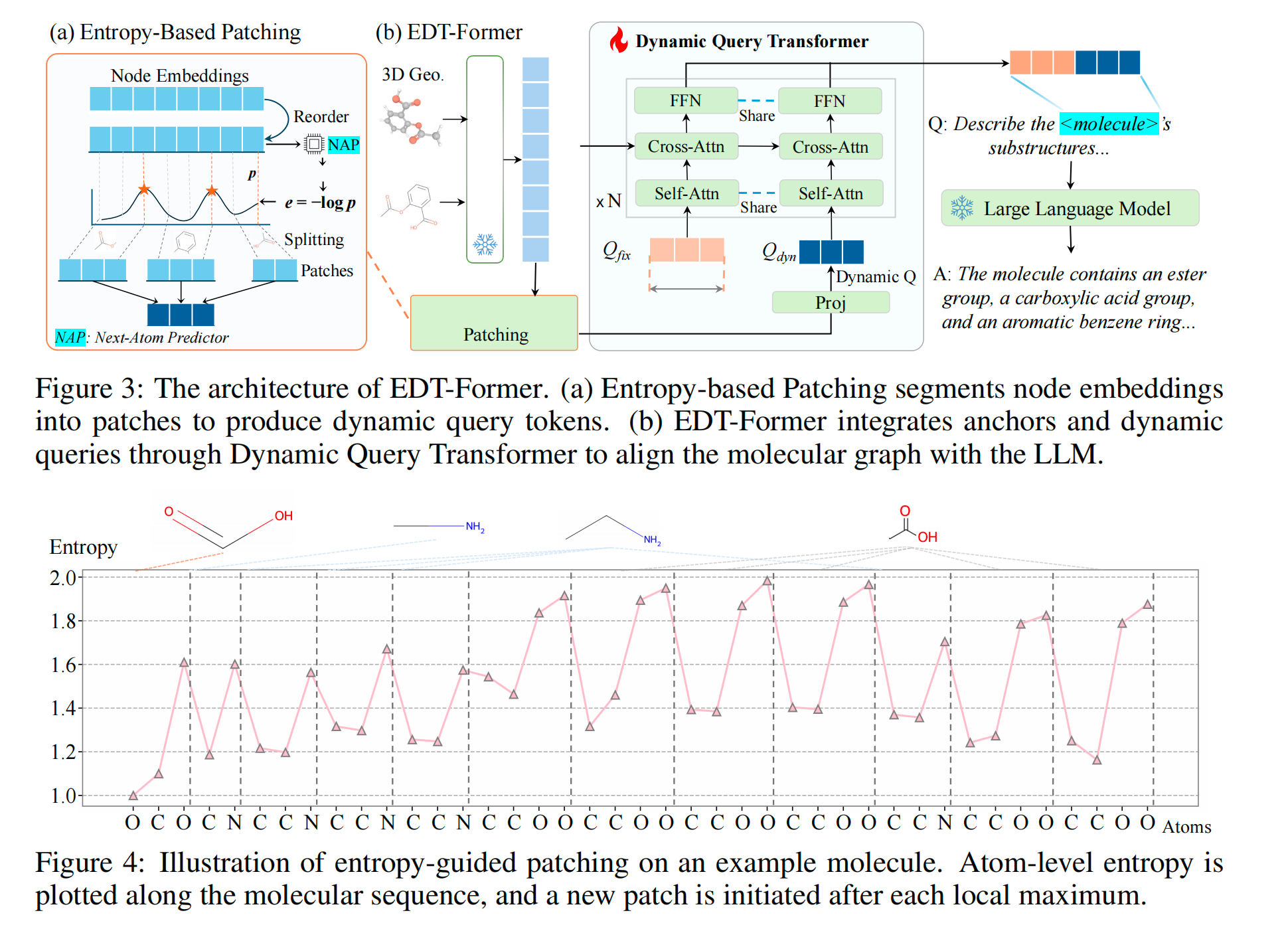
A lightweight predictor, trained to model sequential structure, produces high entropy at positions where the next element is hard to predict — branch points, ring closures, and functional transitions. These entropy peaks become natural split points, dividing the entity into segments. Each segment is pooled into a single token, so more complex entities automatically produce more tokens.
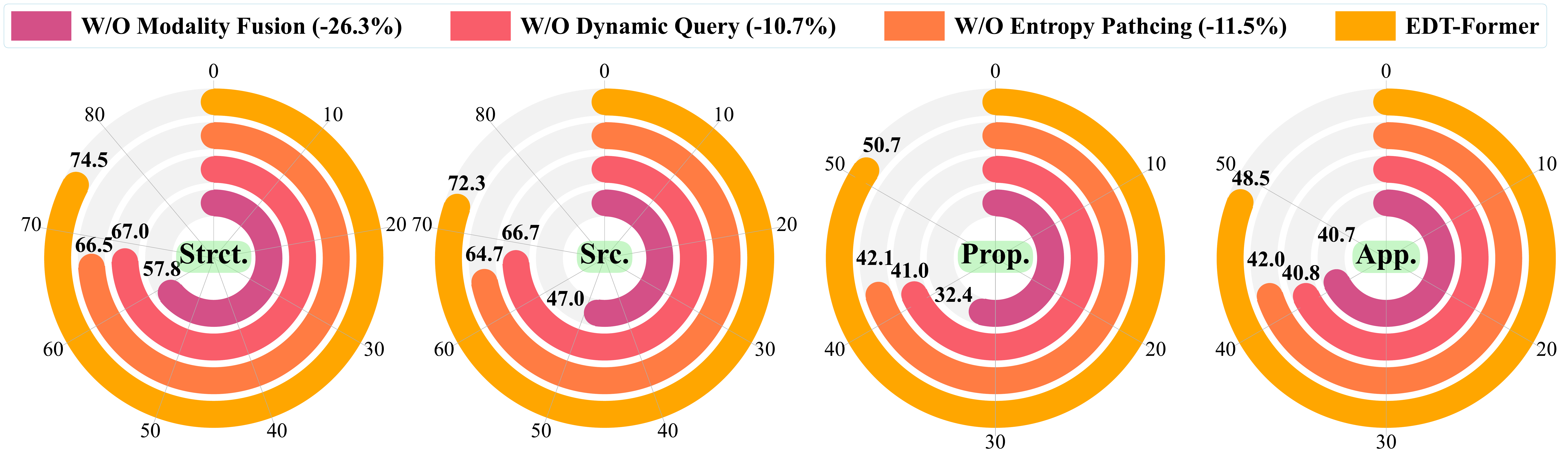
Ablation: replacing learned entropy-based boundaries with uniform fixed-size patches drops average reasoning accuracy by 11.5%.
Adaptive Query Architecture
The dynamic tokens produced above are combined with a set of learned anchor tokens into a shared query pool. A lightweight transformer applies self-attention across this pool to mix global and local context, then cross-attention against the frozen encoder's representations to retrieve detailed structural evidence. Only this connector is trained — both the encoder and LLM remain fully frozen.
Ablation: replacing this architecture with a fixed-length projection drops reasoning accuracy by 10.7%; removing the cross-attention entirely causes a 26% average drop.
Why the Frozen Design Works
Training proceeds in two stages: the connector is first pretrained on structural reconstruction objectives, then aligned to instruction-following with the LLM frozen throughout. Both components contribute independently — removing either one degrades performance across all tasks, and removing multimodal fusion causes the largest single ablation drop.
Results
Empirical Performance
Evaluation across zero-shot property prediction, large-scale reasoning (MoleculeQA), and generation (Mol-Instructions):
| Metric | Result |
|---|---|
| Zero-shot property tasks ranked #1 | 9 / 10 |
| PAMPA accuracy (zero-shot) | 82.34% (+8 pp over best baseline) |
| BBBP accuracy (zero-shot) | 72.48% (+9 pp over second-best) |
| CLINTOX accuracy (zero-shot) | 56.55% (+29 pp over second-best) |
| MoleculeQA avg accuracy (SFT, 8B) | 61.56% — best on all 4 sub-tasks |
| 10-shot EDT-Former vs GPT-5 on MoleculeQA | EDT-Former 8B outperforms GPT-5 |
| Mol-Instructions property MAE | 0.0062 (best; next-best 0.0079) |
| Training time vs LoRA fine-tuning | 3.5× faster per step |
| GPU memory vs LoRA fine-tuning | 37 GB vs 77 GB (2.1× reduction) |
EDT-Former consistently outperforms Mol-LLaMA, 3D-MolM, LLaMo, GPT-4o, and GPT-5 under matched conditions — demonstrating that connector-only frozen-backbone alignment generalizes better than backbone fine-tuning across reasoning, understanding, and generation tasks.
Field Contribution
EDT-Former demonstrates that strong structure-to-language alignment does not require fine-tuning the LLM — a connector that adapts token budgets to entity complexity outperforms systems with full backbone training. The adaptive design is backbone-agnostic and generalizes beyond molecular entities to any structured input with a sequence-ordered representation.
Open-Source Access
EDT-Former is fully open-sourced. All assets are freely available for research and downstream use:
| Asset | Link |
|---|---|
| Paper | arXiv 2602.02742 |
| Code | github.com/zihao-jing/EDT-Former |
| Pretrained Encoder | zihaojing/EDT-Former-encoder |
| Full Model | zihaojing/EDT-Former-model |
| SFT Dataset | zihaojing/EDT-Former-sft-data |
| Pretrain Dataset | zihaojing/EDT-Former-pretrain-data |
Quick Start
conda env create -f environment.yml
conda activate edtformer
pip install --no-deps --no-build-isolation torch-geometric flash-attn torch-scatter
cp env.sh local.env.sh
# Edit local.env.sh: set HF_HOME, BASE_DIR, DATA_DIR, CHECKPOINT_DIR
source local.env.sh
Load Pretrained Model
from huggingface_hub import snapshot_download
snapshot_download("zihaojing/EDT-Former-encoder",
local_dir="checkpoints/edt_former_s1_large/final_model")
snapshot_download("zihaojing/EDT-Former-model",
local_dir="checkpoints/edt_former_s2_large/final_model")
Run Training / Inference
bash scripts/training/pretraining.sh # Stage 1: encoder pretraining
bash scripts/training/finetuning.sh # Stage 2: alignment tuning
bash scripts/qa/mol_qa.sh # MoleculeQA downstream task
bash scripts/qa/mol_qa_scaffold.sh # Scaffold-split variant
For full documentation, dataset layout, DeepSpeed config, and all downstream task scripts, see the GitHub README →
Citation
@inproceedings{jing2026edtformer,
title={Entropy-Guided Dynamic Tokens for Graph-LLM Alignment in Molecular Understanding},
author={Jing, Zihao and Zeng, Qiuhao and Fang, Ruiyi and Sun, Yan and Wang, Boyu and Hu, Pingzhao},
booktitle={International Conference on Learning Representations (ICLR)},
year={2026}
}
Contact
Zihao Jing (first author) — zjing29@uwo.ca
Pingzhao Hu (corresponding author) — phu49@uwo.ca
Questions about EDT-Former, requests to use it as a baseline, or collaboration inquiries are welcome — reach out anytime.