Scaling-Aware Adapter
ICML 2026
ICML 2026LLMMultimodalBiologyAdapter
Overview
Biological sequence data—DNA, RNA, and proteins—exhibits rich multi-scale structure that standard LLM tokenization schemes ignore. Uniform patching treats all positions equally regardless of their information density, causing the model to over-allocate capacity to low-entropy regions and under-represent functionally critical sites.
Scaling-Aware Adapter (EntroAdap) addresses this with two components:
- Scaling-Aware Patching — dynamically adjusts patch granularity based on local entropy, producing denser tokens where sequence complexity is high and coarser tokens in repetitive regions.
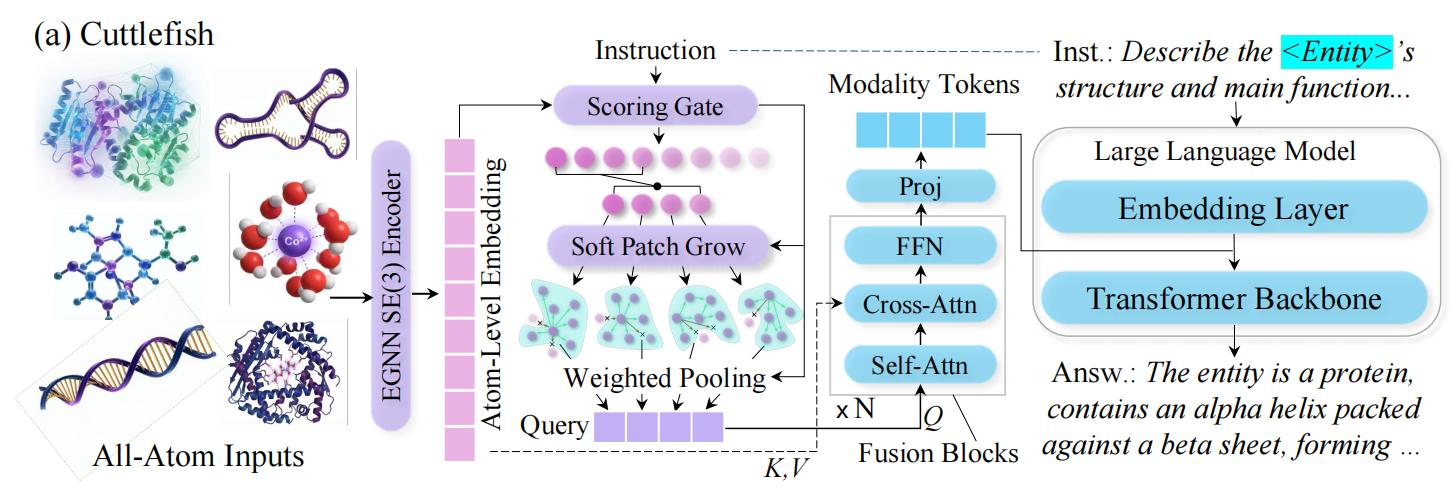
- Geometry-Grounding Adapter — a lightweight connector module that injects 3D structural priors (torsion angles, distance maps) into the LLM's token stream without modifying the backbone weights.
Results
| Benchmark | Tasks Won |
|---|---|
| Mol-Instruction | 7/7 |
| RNA-QA | 5/6 |
| DNA-Chat | 5/5 |
| Overall | 17/18 |
Top-1 performance across all three benchmarks with connector-only training — the LLM backbone is frozen throughout.
Method
The adapter operates in two stages:
- Stage 1 — Entropy Estimation: A lightweight CNN scans the input sequence and produces a per-position entropy score used to compute adaptive patch boundaries.
- Stage 2 — Geometry Injection: Structural descriptors extracted from AlphaFold-predicted coordinates are projected into the LLM's embedding space via a cross-attention adapter layer inserted before the first transformer block.
This design keeps the total trainable parameter count under 5% of the full model, making it efficient for continual adaptation across new biological modalities.