MuMo: Geometry-First Fusion Solves Instability in Multimodal Structure Encoding
NeurIPS 2025
Situation: Multimodal encoders that combine 2D and 3D structural information face two compounding failures — noisy 3D inputs corrupt embeddings when fused directly, and symmetric cross-modal integration lets the noisier stream distort the others. Task: Build a robust multimodal encoder that handles noisy geometric inputs without instability or representation collapse. Action: Built an encoder that first consolidates 3D geometry into a stable, rotation-invariant prior before injecting it into a sequence backbone in a staged, asymmetric way — letting semantics establish before structure modifies them. Result: Ranked #1 on 17 of 22 benchmarks, averaging +2.7% over the best baseline per task, with up to 27% error reduction on conformer-sensitive prediction tasks.
Background & Challenge
Two persistent failure modes in multimodal structural encoding:
1. Noisy 3D inputs destabilize geometric fusion. 3D structural inputs generated by standard tools exhibit high variance for the same entity — different conformers produce conflicting embeddings, making structurally similar entities indistinguishable in embedding space. This noise propagates through naive fusion architectures, corrupting downstream predictions.
2. Symmetric fusion causes modality collapse. When all modalities are treated as equally clean and fused uniformly, the noisier 3D stream distorts the more stable 2D and sequence representations. Existing models that concatenate or uniformly attend across modalities suffer degraded representation quality as a result.
Methodology
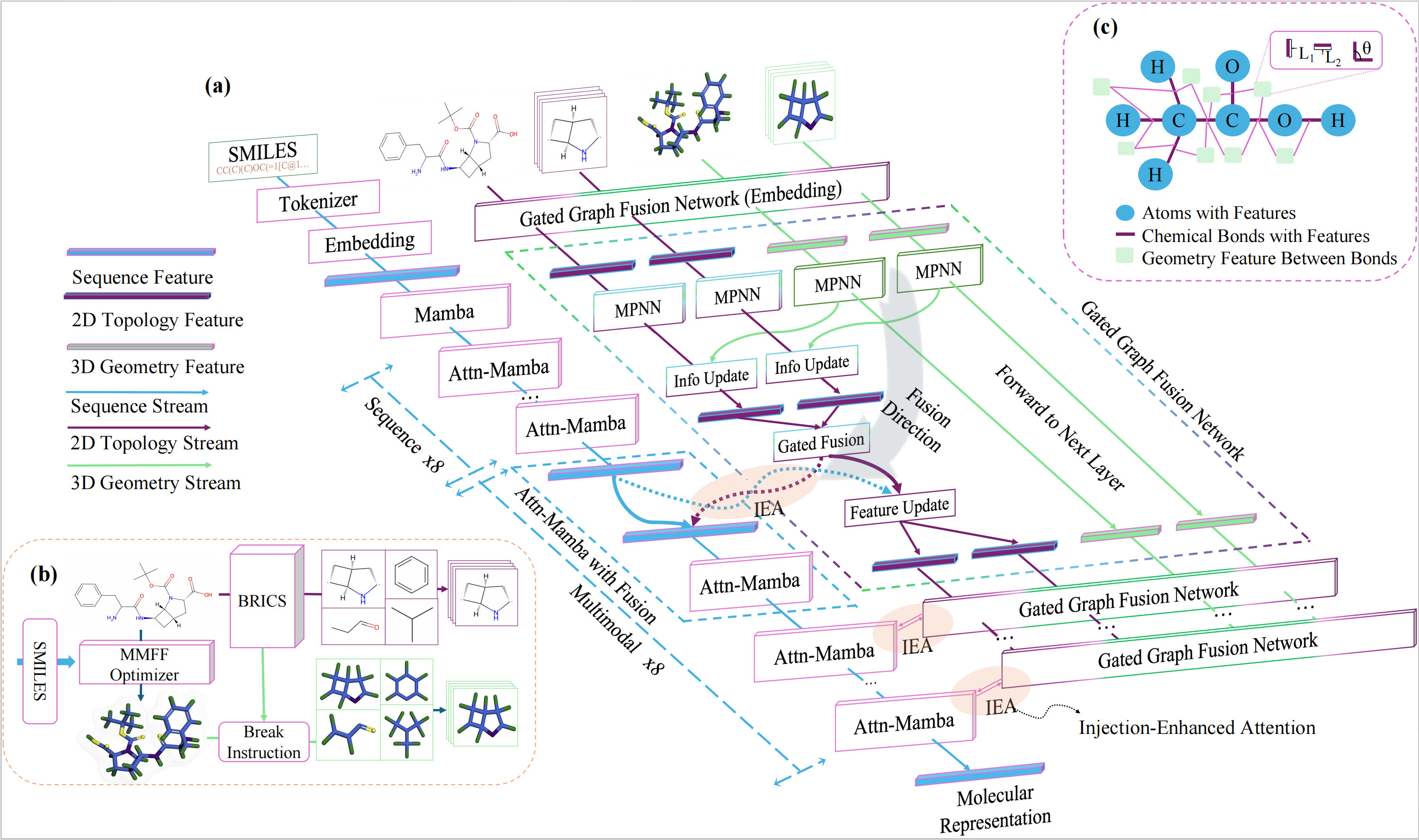
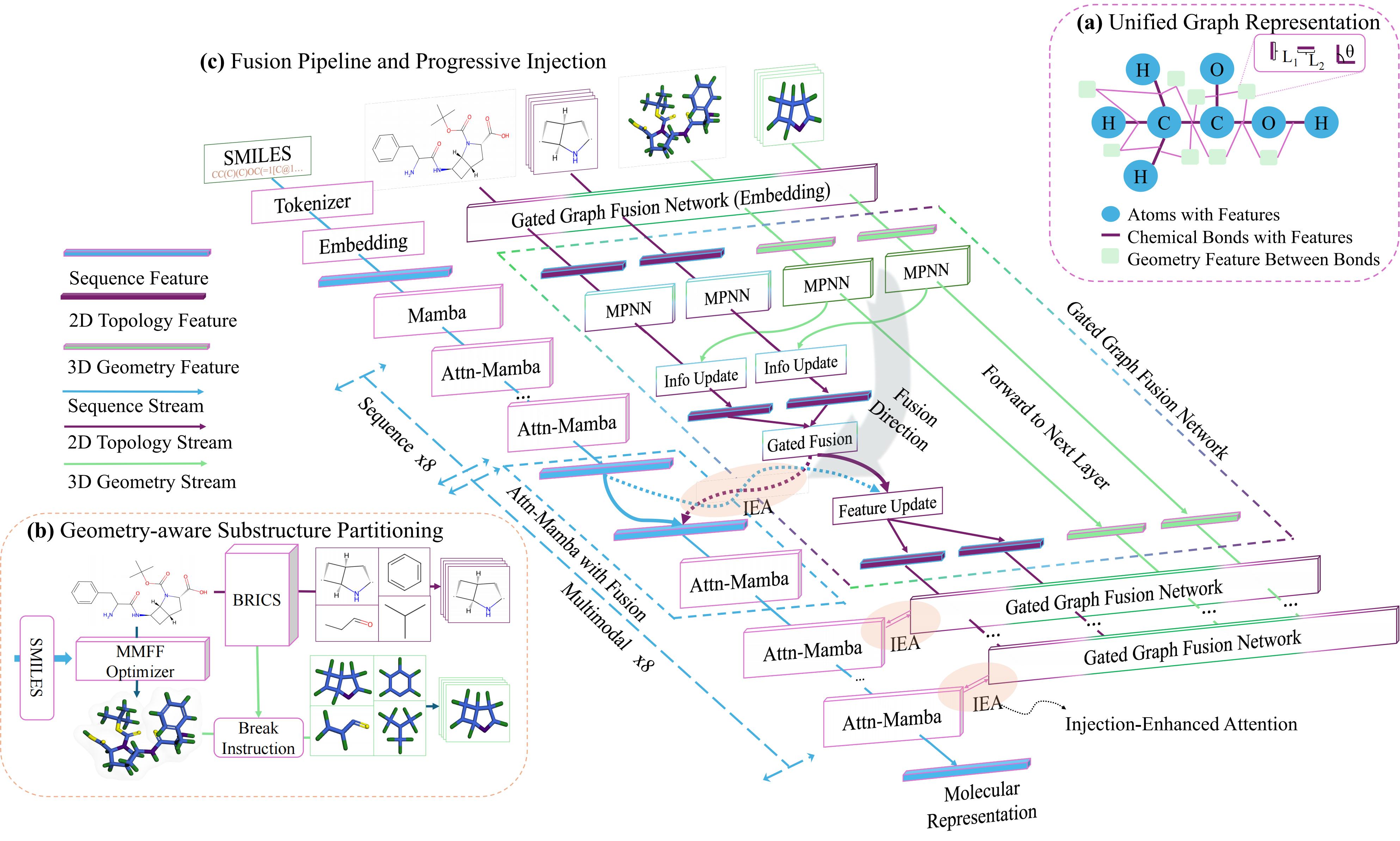
Geometry Stabilization
Before any cross-modal fusion, we first consolidate the 3D geometry into a stable, rotation-invariant prior. Each entity is represented as a unified graph that encodes both 2D bond connectivity and 3D spatial constraints. Message passing propagates both signals jointly, and global and local structural views are combined into a single prior that is robust to conformer variation — removing the noise before it has any chance to corrupt the sequence stream.
Asymmetric Staged Injection
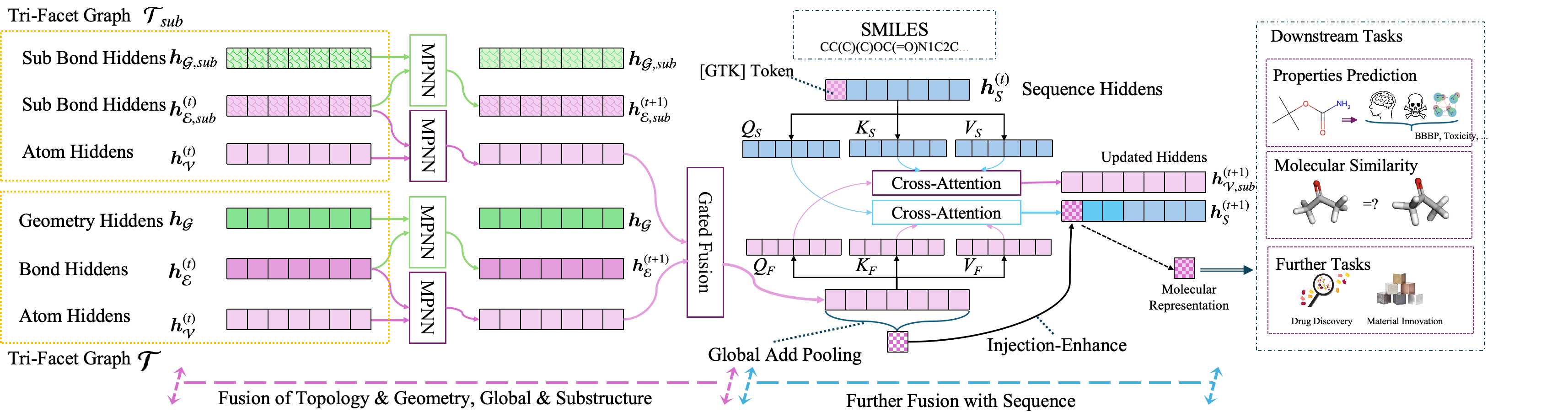
The stable structural prior is injected into the sequence backbone only in the last half of layers, letting the model first build semantic context from the sequence before introducing geometric constraints. Cross-attention aligns the structural prior with the sequence at each injection point, and a pooled global representation is added to a dedicated anchor token via a residual update.
Ablation: late injection alone gives +14.5%; our staged timing gives +17.1% — a 2.6 pp gain from injection timing alone.
Why Multimodal Training Works
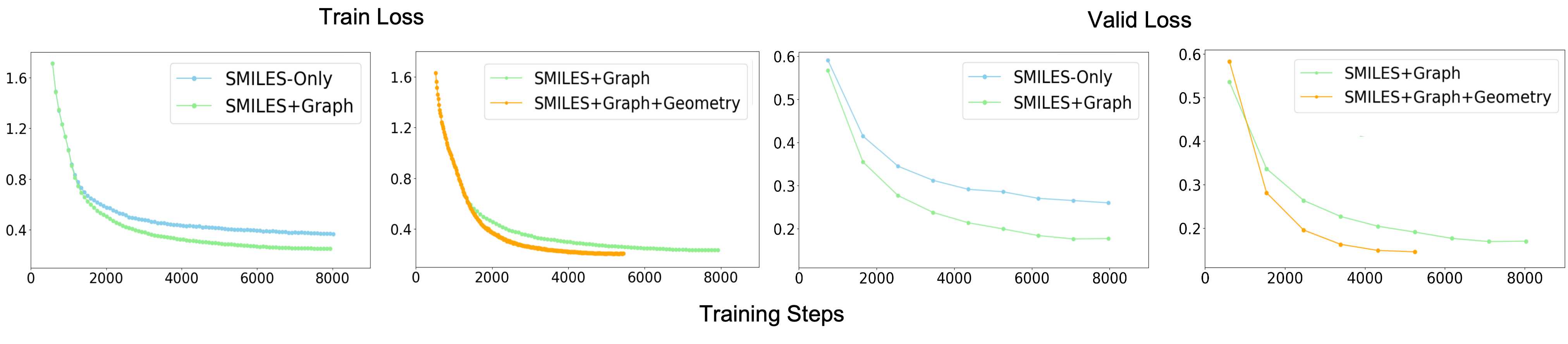
Pretraining loss curves directly validate the fusion design: each modality contributes a non-redundant signal — adding 2D topology accelerates convergence, and further adding 3D geometry achieves the lowest final loss. Unimodal baselines converge to a higher floor on every objective.
Results
Empirical Performance
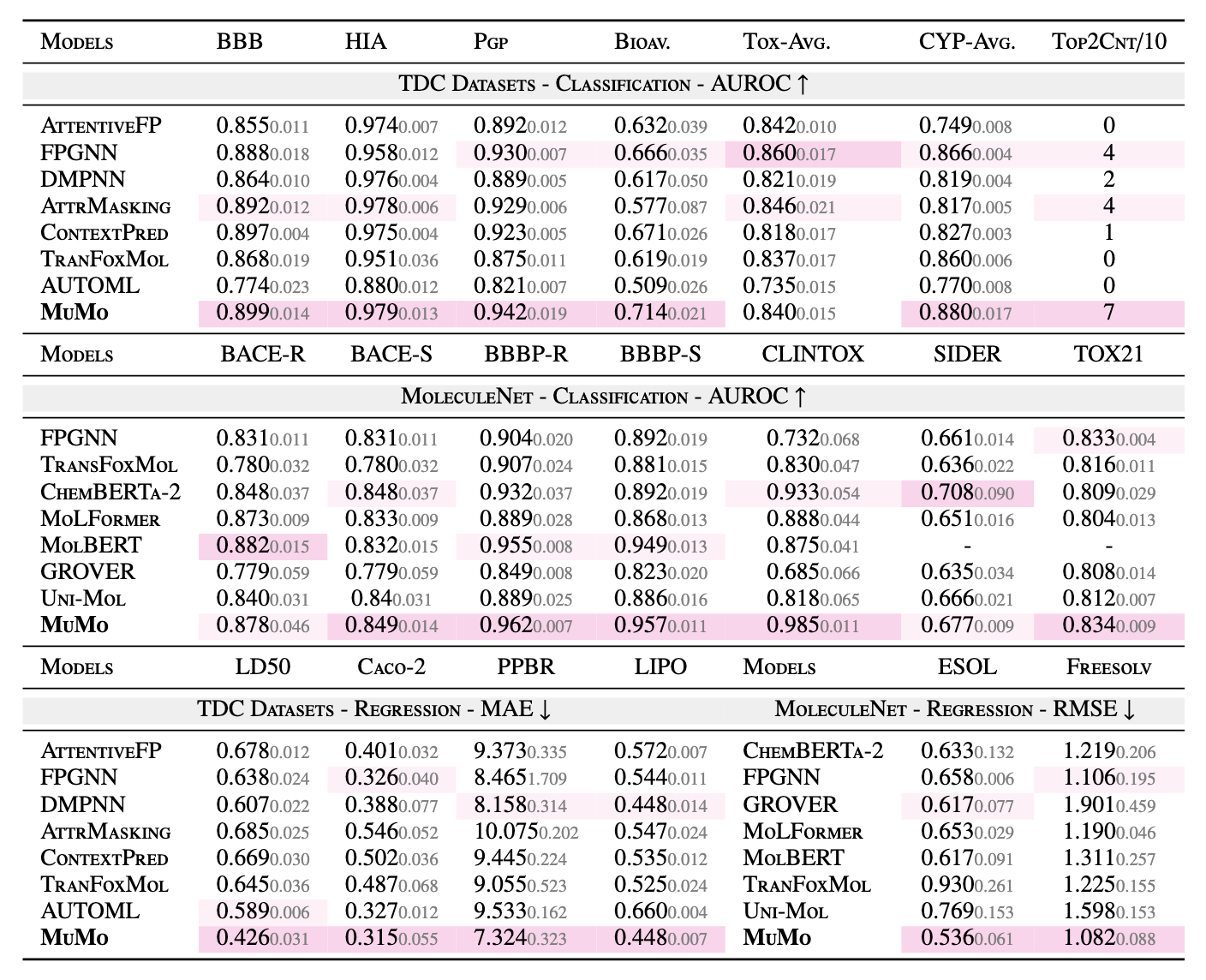
Comprehensive evaluation across 22 benchmark tasks (TDC ADMET + MoleculeNet), covering both classification and regression:
| Metric | Result |
|---|---|
| Tasks ranked #1 | 17 / 22 |
| Avg. improvement over best baseline per task | +2.7% |
| Best single-task gain (LD50 toxicity) | 27% MAE reduction |
| Conformer-sensitive regression (PPBR, LD50) | Lowest error among all models |
| Broader benchmarks (Reaxtica) | 3 / 4 tasks ranked #1 |
| Ablation: removing 3D geometry | −7.46% avg |
| Ablation: removing sequence only | −13.11% avg |
MuMo consistently outperforms Uni-Mol, GROVER, MoLFormer, and ChemBERTa-2 — demonstrating that structured, asymmetric fusion generalizes better than both unimodal encoders and symmetric multimodal architectures.
Field Contribution
MuMo addresses two failure modes — geometry instability and modality collapse — that prior encoders treated as separate unsolved problems, closing both through precise architectural choices rather than scale. The design is backbone-agnostic and generalizes to any domain with heterogeneous structured inputs; it has been validated at scale through large-scale virtual screening, confirming that robust multimodal fusion transfers directly to real deployment workloads.
Open-Source Access
MuMo is fully open-sourced. All assets are freely available for research and downstream use:
| Asset | Link |
|---|---|
| Paper | NeurIPS 2025 (arXiv 2510.23640) |
| Code | github.com/zihao-jing/MuMo |
| Pretrained Model | zihaojing/MuMo-Pretrained |
| Finetuning Datasets | zihaojing/MuMo-Finetuning |
| Pretraining Dataset | zihaojing/MuMo-Pretraining |
Quick Start
# Clone and set up environment
git clone https://github.com/zihao-jing/MuMo.git
conda env create -f environment.yml && conda activate mumo
# Configure paths (edit BASE_DIR and DATA_DIR)
nano init_env.sh
source init_env.sh
Load Pretrained Model
from transformers import AutoConfig, AutoTokenizer
from model.load_model import load_model
from dataclasses import dataclass
repo = "zihaojing/MuMo-Pretrained"
config = AutoConfig.from_pretrained(repo, trust_remote_code=True)
tokenizer = AutoTokenizer.from_pretrained(repo)
@dataclass
class ModelArgs:
model_name_or_path: str = repo
model_class: str = "MuMoFinetune" # or "MuMoPretrain"
task_type: str = "classification" # or "regression"
cache_dir: str = None
model_revision: str = "main"
use_auth_token: bool = False
model = load_model(config, tokenizer=tokenizer, model_args=ModelArgs())
Finetune & Inference
# Finetune on TDC regression (LD50 toxicity)
bash scripts/sft_tdc/regression/LD50.sh
# Batch inference / virtual screening
bash scripts/infer/infer_ic50.sh
For the full documentation, dataset layout, DeepSpeed training setup, and reproducibility instructions, see the GitHub README →
Citation
@inproceedings{jing2025mumo,
title = {MuMo: Multimodal Molecular Representation Learning via
Structural Fusion and Progressive Injection},
author = {Jing, Zihao and Sun, Yan and Li, Yan Yi and
Janarthanan, Sugitha and Deng, Alana and Hu, Pingzhao},
booktitle = {Advances in Neural Information Processing Systems (NeurIPS)},
year = {2025}
}
Contact
Zihao Jing (first author) — zjing29@uwo.ca
Pingzhao Hu (corresponding author) — phu49@uwo.ca
Questions, collaborations, or requests to use MuMo as a baseline? We're happy to help — reach out anytime.